forge-db: SIMD-Accelerated Vector Search
High-performance vector database with IVF-PQ and HNSW indexing. Pure Rust, no external dependencies.
Vector search powers semantic search, RAG retrieval, recommendation systems, and duplicate detection by finding the k nearest neighbors to a query embedding. The naive approach computes distances to every vector in the database, which guarantees 100% recall but scales linearly with dataset size. When searching millions of vectors, this becomes prohibitively expensive. Approximate nearest neighbor algorithms trade small recall losses for orders of magnitude speedups by avoiding exhaustive comparison.
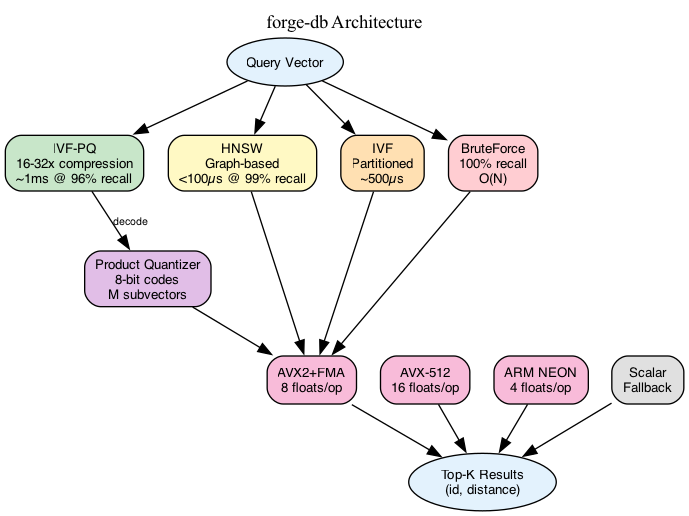
Forge-db implements multiple indexing strategies with different performance characteristics. Brute force serves as the correctness baseline and works well for small datasets. IVF partitions the vector space using k-means clustering and searches only the nearest partitions at query time. IVF-PQ adds product quantization to compress vectors from 3KB to 32 bytes while using lookup tables for fast approximate distance computation. HNSW builds hierarchical proximity graphs for logarithmic search but requires significantly more memory. Users choose based on their latency, recall, and memory constraints.
Product quantization compresses vectors by splitting each into subvectors and quantizing them independently against learned codebooks. A 768-dimensional float32 vector normally requires 3KB. PQ splits this into 32 subvectors of 24 dimensions each, quantizes each to the nearest of 256 centroids, and stores the result as a single byte per subvector. The compressed representation is 32 bytes, a 96x reduction. Distance computation becomes table lookups instead of floating point operations. The query vector’s distance to all centroids is precomputed once, then database vectors are scored by summing the distances corresponding to their codes.
The accuracy tradeoff is concrete. With nprobe=16 clusters and no reranking, IVF-PQ achieves 69% recall at 520 microseconds per query using 31MB of memory. Adding a reranking pass where the top 100 compressed results are rescored using full precision improves recall to 96% at 1 millisecond latency. The memory footprint stays constant because full-precision vectors are loaded only for the reranking candidates. This two-stage approach recovers most of the quantization error while maintaining the memory benefits.
SIMD and Performance Optimization
Distance computation dominates query latency. Computing cosine similarity for a 768-dimensional vector requires 768 multiply-add operations per comparison. SIMD instructions process multiple floats per instruction, providing linear speedups. The implementation detects CPU features at runtime and dispatches to the optimal code path. AVX-512 processes 16 floats per instruction, AVX2 with FMA processes 8 floats, ARM NEON handles 4 floats, and a scalar fallback ensures portability. Runtime dispatch means a single binary runs optimally across different processors without compile-time feature flags.
Cache alignment proved unexpectedly impactful on performance. Aligning vector data to 64-byte boundaries improved throughput by 15% by eliminating unaligned SIMD loads. False sharing between threads during parallel index construction caused another 10% performance degradation. Padding shared data structures to cache line boundaries eliminated the false sharing. These optimizations required profiling with performance counters to identify the bottlenecks, which are invisible in algorithmic complexity analysis.
Testing approximate algorithms requires statistical validation. Unlike exact algorithms where correctness is binary, approximate nearest neighbor methods trade accuracy for speed. A recall of 96% means 4% of results differ from ground truth. The test harness compares every query against brute force and flags statistically significant recall regressions. This catches subtle bugs in quantization or distance computation that would otherwise manifest as gradual quality degradation.
Index construction uses k-means clustering for IVF partitioning and codebook learning for product quantization. Both are iterative algorithms that converge over multiple passes. The implementation parallelizes distance computation across threads but carefully manages memory allocation to avoid false sharing. Building an index for a million 768-dimensional vectors takes approximately one minute on a modern CPU. This one-time cost is acceptable because the index is reused for many queries.
Current limitations point toward clear next steps. GPU acceleration would provide another 10-100x speedup for distance computation because the operation is embarrassingly parallel. A CUDA backend could process thousands of distance calculations simultaneously. Binary quantization using 1-bit codes and Hamming distance offers even faster candidate filtering than product quantization, though with different accuracy tradeoffs. This would enable a three-tier system: binary quantization for initial filtering, PQ for approximate scoring, and full precision for final reranking.
The codebase uses pure Rust with no external dependencies for the core algorithms. This simplifies deployment and ensures reproducible builds. SIMD intrinsics require unsafe blocks but are isolated and tested exhaustively. The API exposes index configuration through a builder pattern, letting users adjust parameters like cluster count, codebook size, and construction threads without recompiling.
Working on forge-db demonstrated that performance optimization requires measurement at every step. Algorithmic improvements like approximate search provide order-of-magnitude gains, but hardware-specific optimizations like cache alignment and SIMD add critical percentage improvements that compound. Testing approximate algorithms demands statistical rigor beyond standard unit tests. Recall can degrade silently through subtle bugs in distance computation or quantization. Real-time feature detection enables shipping a single binary that performs optimally across processors, which matters more in practice than compile-time specialization.